About Bionamic
Founding
Bionamic, based in Lund, Sweden, was founded in 2019 by experts in life science, theoretical physics, and software development with the goal of providing the best platform in the world for antibody research.
The idea behind Bionamic was developed while consulting at various biopharma companies, where the challenges of organizing and analyzing development data were obvious and researchers were frustrated by the inefficiencies caused by the lack of appropriate software.
Founders

Anders Carlsson, Ph.D.
"Our journey with Bionamic began with a simple idea: to develop a software solution that could support the antibody discovery and development process. We understood the frustrations that researchers faced when organizing and analyzing development data, and we are committed to providing a software solution that is built on true biological concepts and relations."

Morten Krogh, Ph.D.
"At Bionamic, we don't believe in imposing rigid workflows on our customers. Instead, our platform is designed to be flexible and adaptable, empowering users to work the way they want to work. We understand that every Bio-Pharma company is unique, and we are committed to providing a solution that fit their specific needs."
The Problem
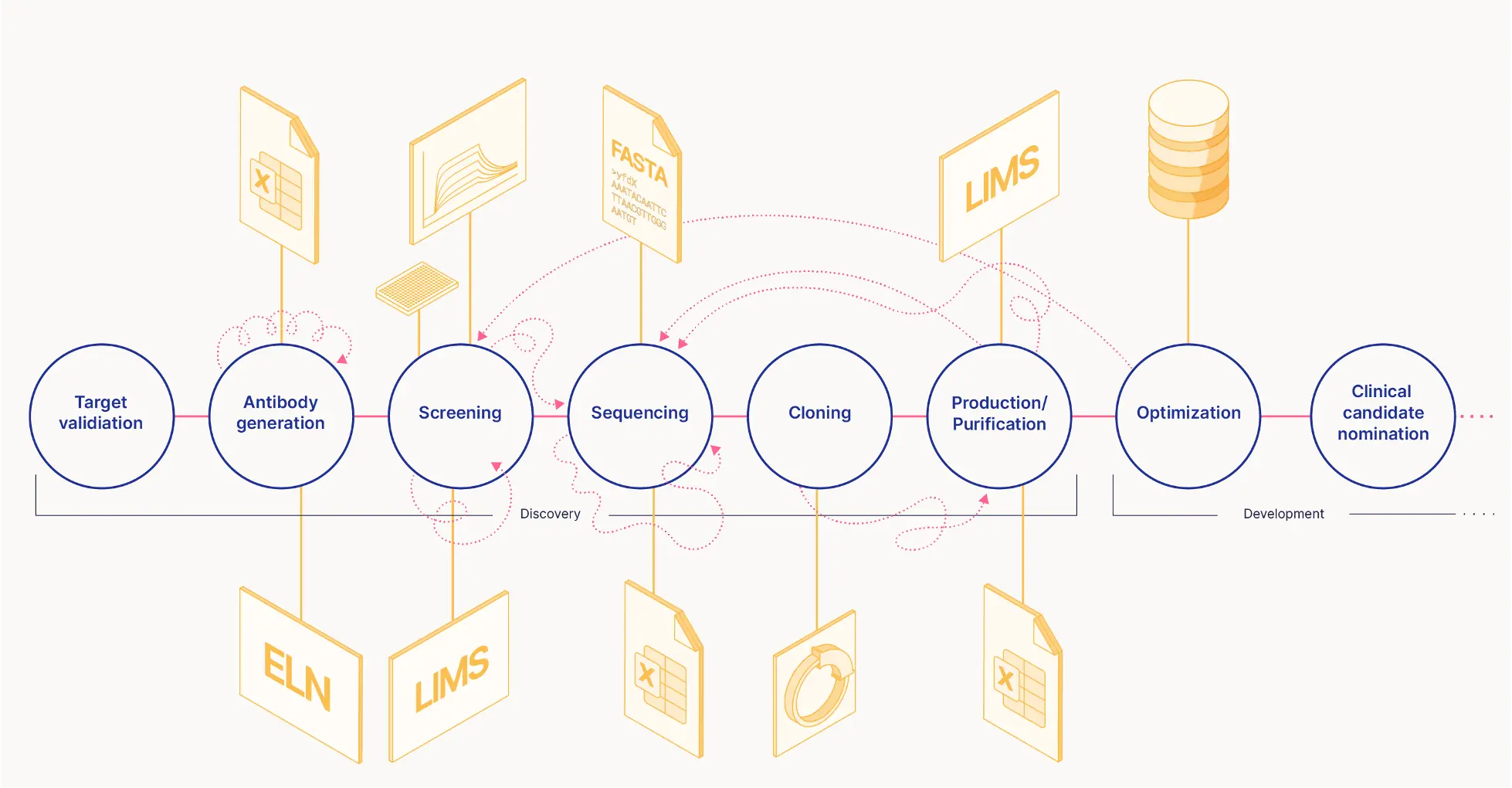
Digitalizing biopharma research has been slow. Current software is often designed for more standardized workflows and solves a specific problem. The discovery and development data is spread out across many files and systems. When employees leave, data is often orphaned. It is too difficult for life-scientists to interact with the systems.
Our Solution
The Bionamic platform keeps track of all antibody discovery and development data in one system. Life scientists interact with the system through a simple UI. The Bionamic platform also allows for advanced scripting that allows bioinformaticians to automate workflows. The Bionamic platform combines data storage with statistics, visualization, sequence alignment and other tasks giving a holistic view of all data.